Lecture 14: Reducing Items and Attributes
Brian J. Smith
2026-03-26
Reducing Items and Attributes
Reducing Items and Attributes
Why?
- Manage complexity in our visualizations.
- Find a strategy that reduces the visual complexity while minimizing the changes of hiding important information from the user.
Reducing Items and Attributes
How?
- Filter:
- Eliminating elements (items or attributes).
- Easy to understand and compute.
- But, “out of sight, out of mind.”
- Aggregate:
- Creating a new single element to replace multiple others.
- Challenging to design and convey.
- Safer cognitively to avoid forgetting filtered information.
Filtering
- Straightfoward and intuitive.
- Just don’t show some things!
- Common when exploring a new dataset.
- Plot only some attributes (columns).
- Often based on knowledge of what the attributes represent.
- Plot only some items (rows).
- Often based on values.
- Sometimes based on a random subset.
Filtering
Attribute Filtering
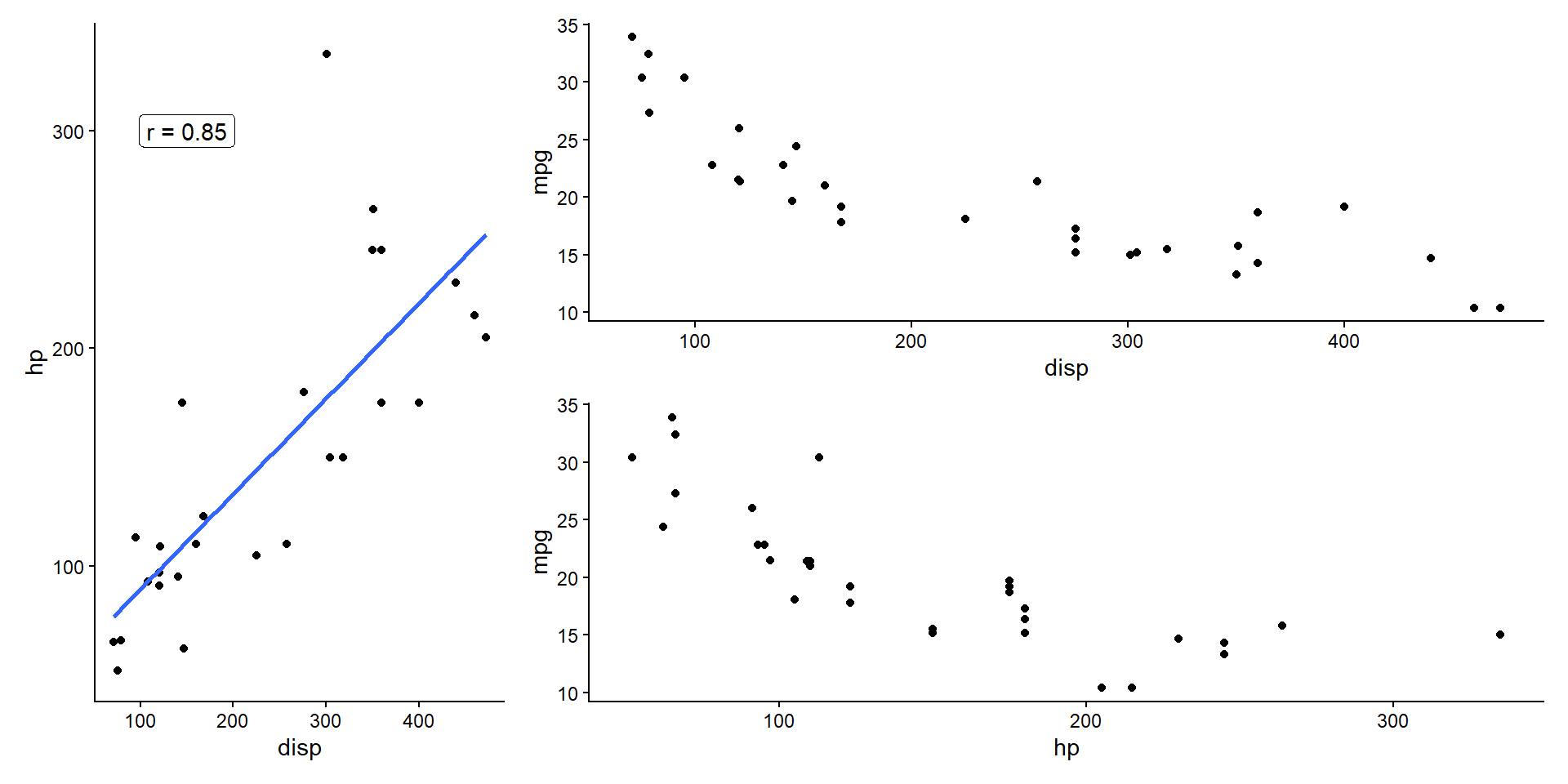
- Often, we have an idea which attributes are most important to us.
- Prioritize visualizations with those attributes.
Filtering
Attribute Filtering
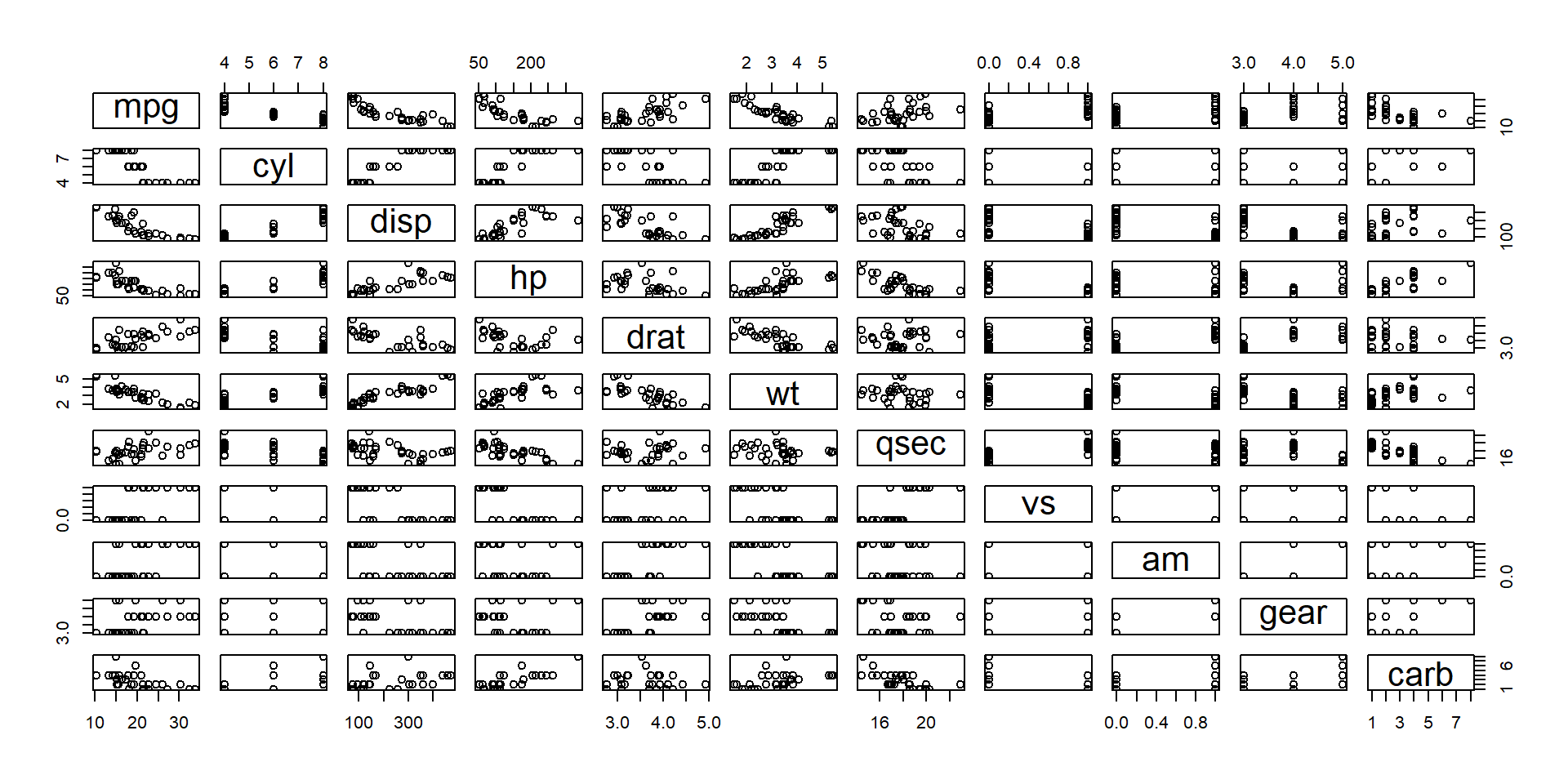
- This may be combined with attribute ordering.
- Orders attributes based on their similarity.
- E.g., highly correlated attributes show some redundant information.
Code
a <- mtcars %>%
ggplot(aes(x = disp, y = hp)) +
geom_point() +
geom_label(x = 150, y = 300,
label = paste("r =",
round(cor(mtcars$disp,
mtcars$hp,
method = "spearman"),
2))) +
geom_smooth(method = "lm", se = FALSE) +
theme_classic()
b <- mtcars %>%
ggplot(aes(x = disp, y = mpg)) +
geom_point() +
theme_classic()
c <- mtcars %>%
ggplot(aes(x = hp, y = mpg)) +
geom_point() +
theme_classic()
des <- "AB
AC"
a + b + c + plot_layout(widths = c(0.3, 0.7),
design = des)Filtering
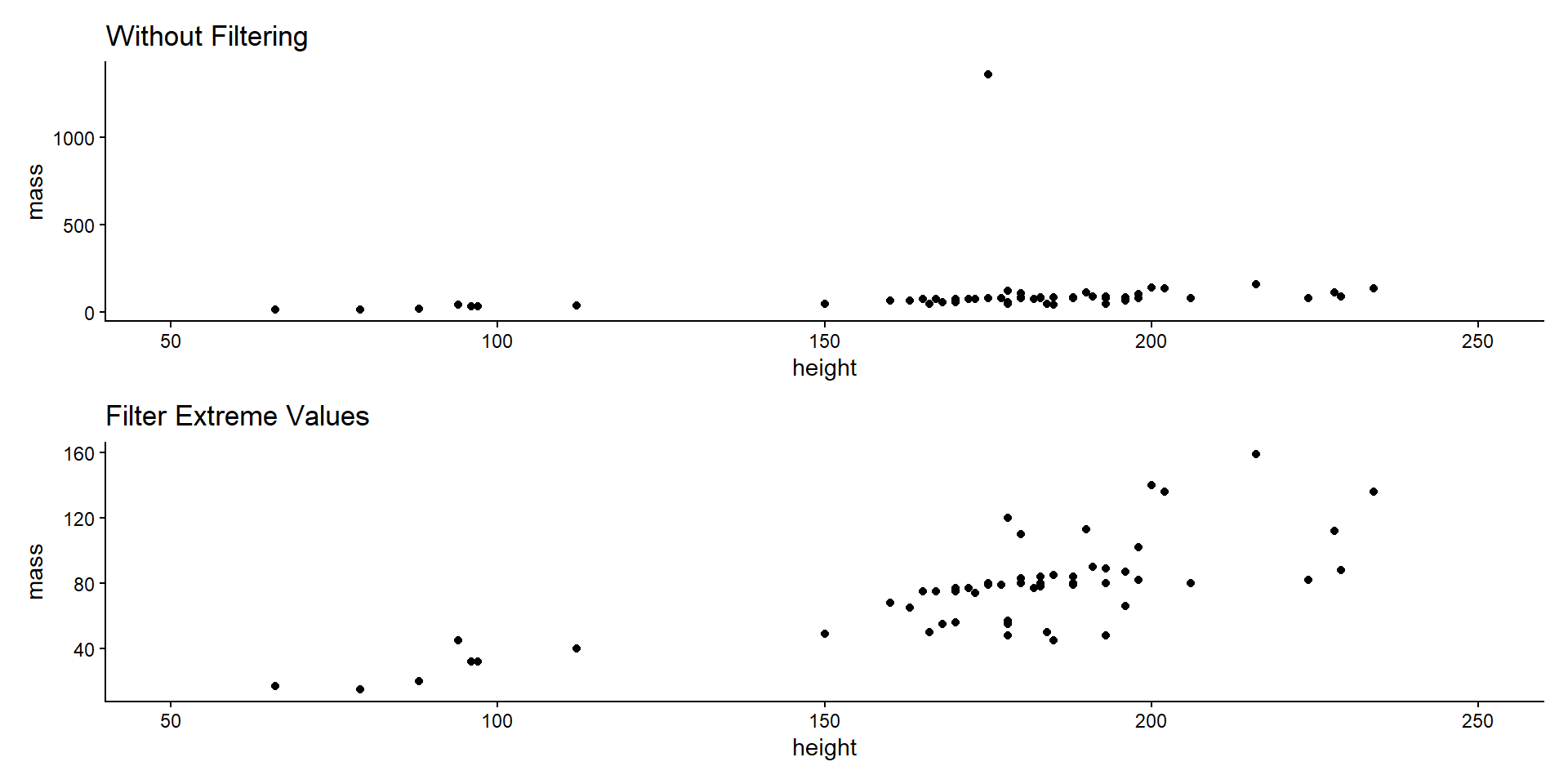
Item Filtering
- Filter items based on values.
- E.g., remove outliers or extreme values.
Code
data("starwars")
a <- starwars %>%
ggplot(aes(x = height, y = mass)) +
geom_point() +
coord_cartesian(xlim = c(50, 250)) +
ggtitle("Without Filtering") +
theme_classic()
b <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(x = height, y = mass)) +
geom_point() +
coord_cartesian(xlim = c(50, 250)) +
ggtitle("Filter Extreme Values") +
theme_classic()
a/bFiltering
Item Filtering
- Randomly subset items.
- Useful for (truly) big datasets where plotting all items would be difficult (impossible).
Aggregation
- Creating a new derived element to represent the group.
- Simple examples include summary statistics.
- Remember Anscombe’s Quartet!
Aggregation
Item Aggregation
- Easily accomplished with idioms that:
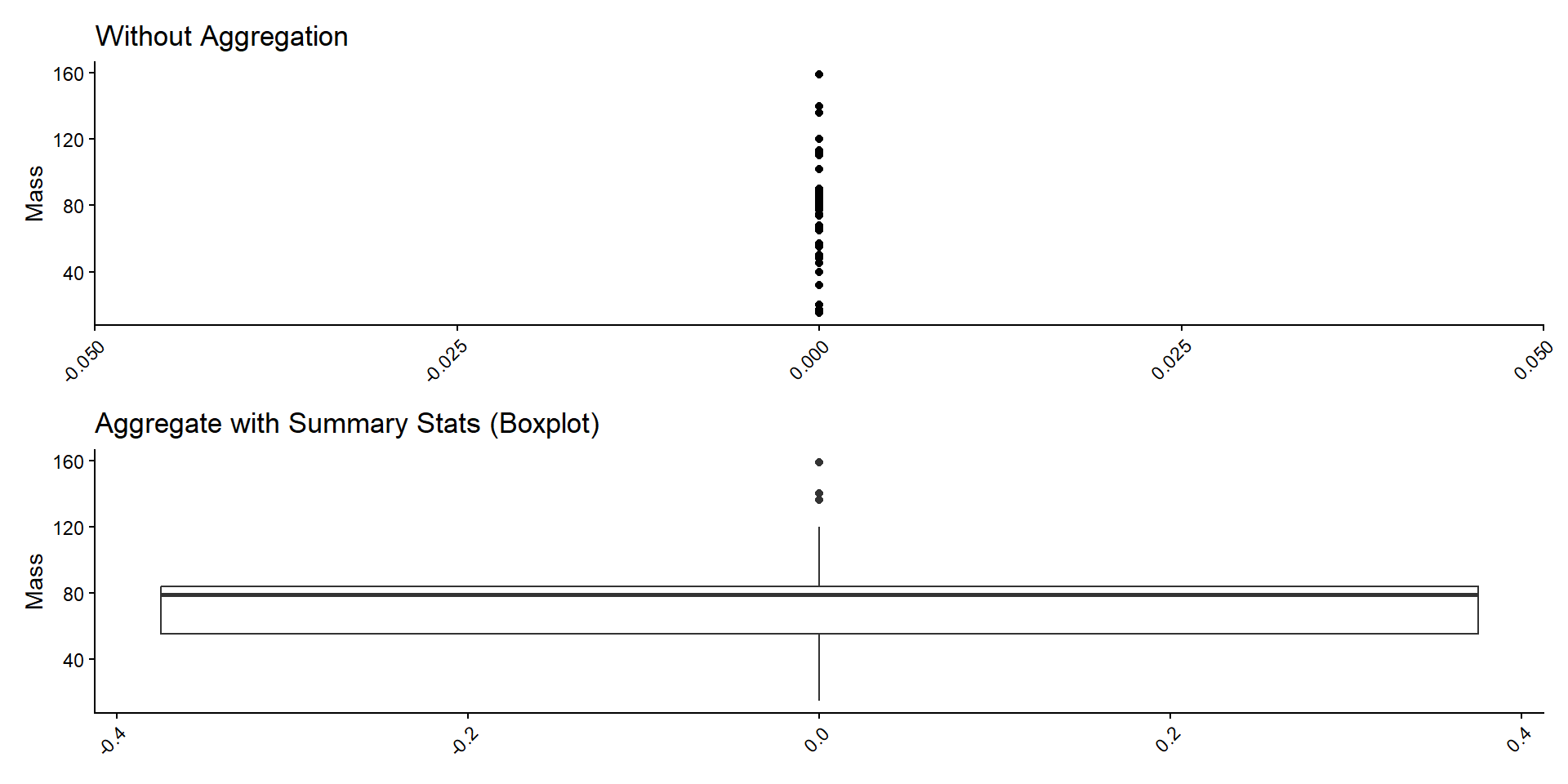
- Display summary statistics.
- Mean, SD, etc. (moments)
- Counts, densities
- Show distributions.
- Display summary statistics.
Aggregation
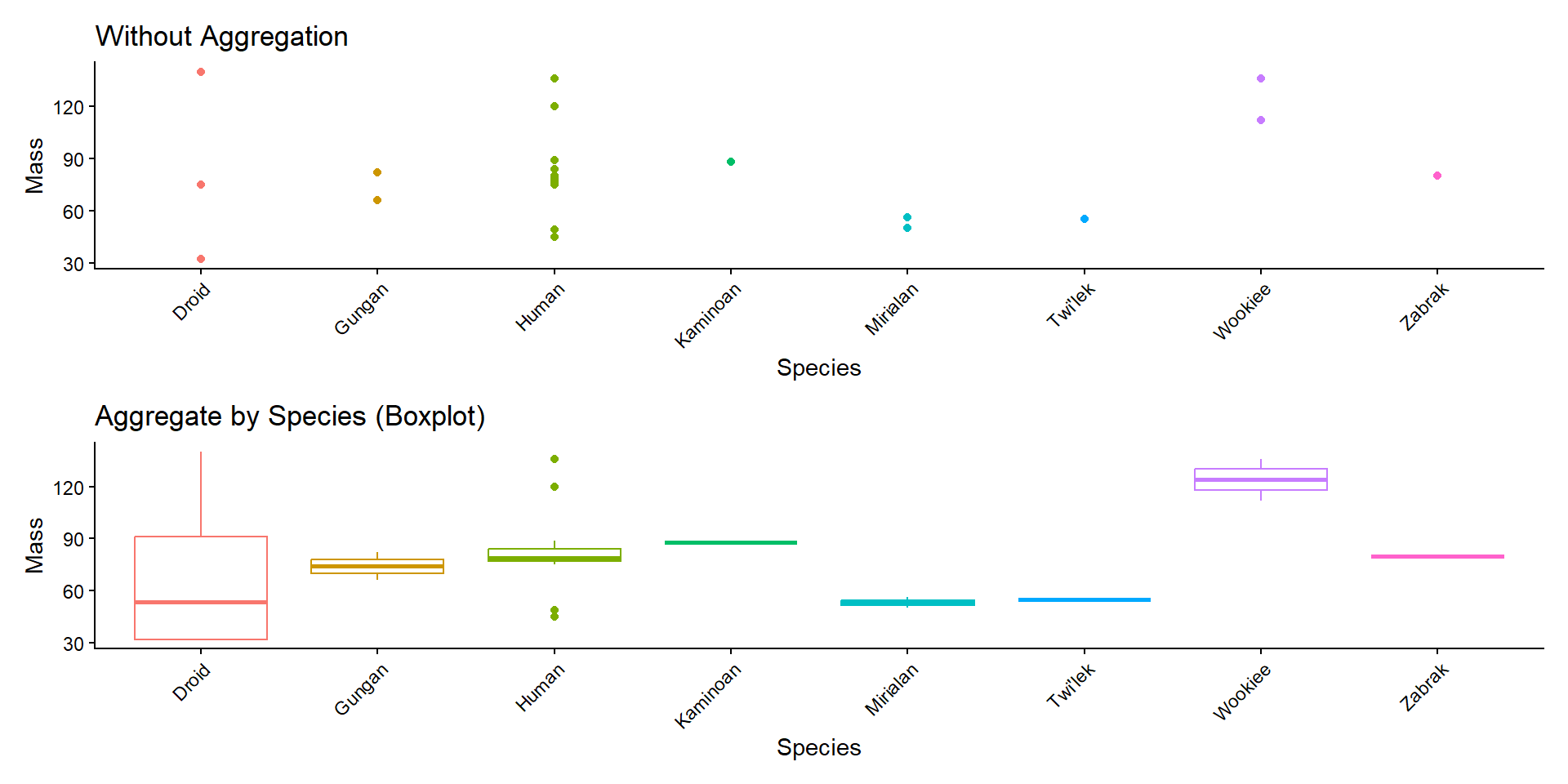
Item Aggregation
1D Summary Stats
Code
a <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(x = 0, y = mass)) +
geom_point(show.legend = FALSE) +
ggtitle("Without Aggregation") +
labs(x = NULL, y = "Mass") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
b <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(y = mass)) +
geom_boxplot(show.legend = FALSE) +
ggtitle("Aggregate with Summary Stats (Boxplot)") +
labs(x = NULL, y = "Mass") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
a/bAggregation
Item Aggregation
2D Summary Stats
Code
spp <- starwars %>%
filter(!is.na(species)) %>%
count(species) %>%
filter(n > 1)
sw <- starwars %>%
filter(species %in% spp$species) %>%
mutate(species = factor(species))
a <- sw %>%
ggplot(aes(x = species, y = mass, color = species)) +
geom_point(show.legend = FALSE) +
ggtitle("Without Aggregation") +
labs(x = "Species", y = "Mass") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
b <- sw %>%
ggplot(aes(x = species, y = mass,
color = species)) +
geom_boxplot(show.legend = FALSE) +
ggtitle("Aggregate by Species (Boxplot)") +
labs(x = "Species", y = "Mass") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
a/bAggregation
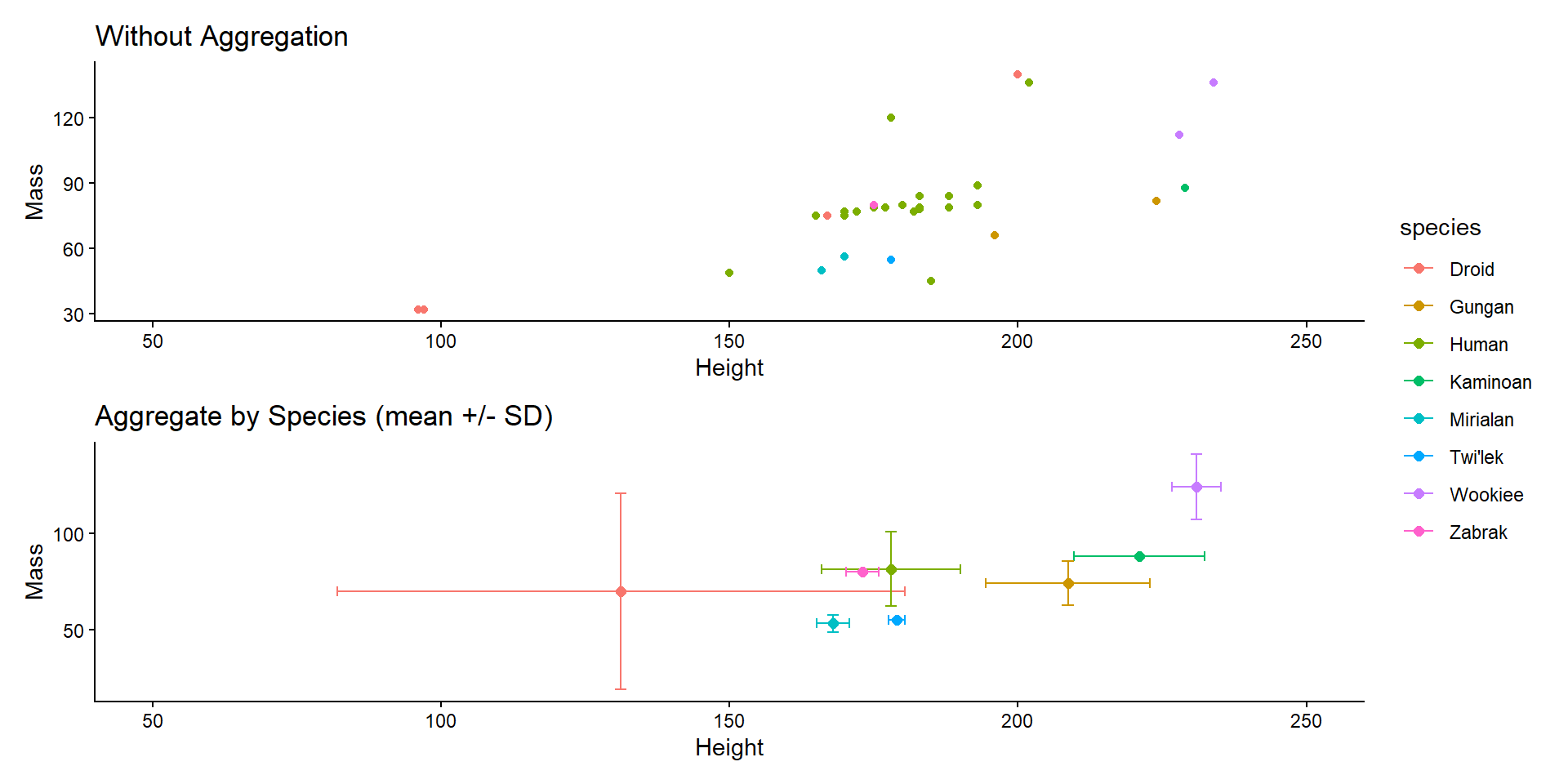
Item Aggregation
3D Summary Stats
Code
# 'sw' created on previous slide
a <- sw %>%
ggplot(aes(x = height, y = mass, color = species)) +
geom_point(show.legend = FALSE) +
coord_cartesian(xlim = c(50, 250)) +
ggtitle("Without Aggregation") +
labs(x = "Height", y = "Mass") +
theme_classic()
b <- sw %>%
group_by(species) %>%
summarize(mass_mean = mean(mass, na.rm = TRUE),
mass_sd = sd(mass, na.rm = TRUE),
height_mean = mean(height, na.rm = TRUE),
height_sd = sd(height, na.rm = TRUE)) %>%
mutate(mass_lwr = mass_mean - mass_sd,
mass_upr = mass_mean + mass_sd,
height_lwr = height_mean - height_sd,
height_upr = height_mean + height_sd) %>%
ggplot(aes(x = height_mean, y = mass_mean,
color = species)) +
geom_errorbar(aes(xmin = height_lwr, xmax = height_upr),
width = 5) +
geom_errorbar(aes(ymin = mass_lwr, ymax = mass_upr),
width = 2) +
geom_point(size = 2) +
coord_cartesian(xlim = c(50, 250)) +
ggtitle("Aggregate by Species (mean +/- SD)") +
labs(x = "Height", y = "Mass") +
theme_classic()
a/b + plot_layout(guides = "collect")Aggregation
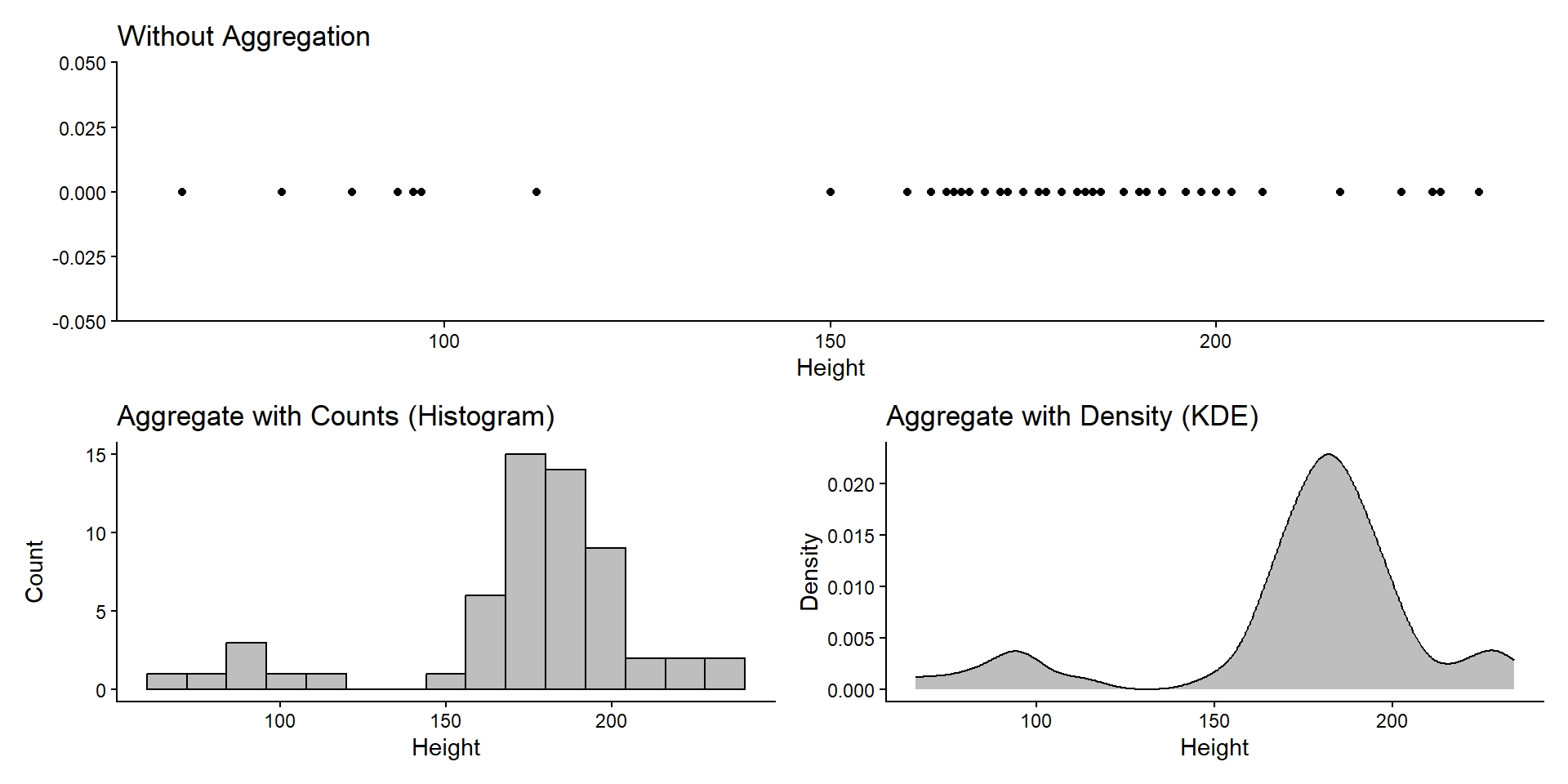
Item Aggregation
1D Counts/Densities
Code
a <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(y = 0, x = height)) +
geom_point(show.legend = FALSE) +
ggtitle("Without Aggregation") +
labs(x = "Height", y = NULL) +
theme_classic()
b <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(x = height)) +
geom_histogram(bins = 15, color = "black", fill = "gray") +
ggtitle("Aggregate with Counts (Histogram)") +
labs(x = "Height", y = "Count") +
theme_classic()
c <- starwars %>%
filter(mass < 500) %>%
ggplot(aes(x = height)) +
geom_density(color = "black", fill = "gray") +
ggtitle("Aggregate with Density (KDE)") +
labs(x = "Height", y = "Density") +
theme_classic()
a/(b+c)Aggregation
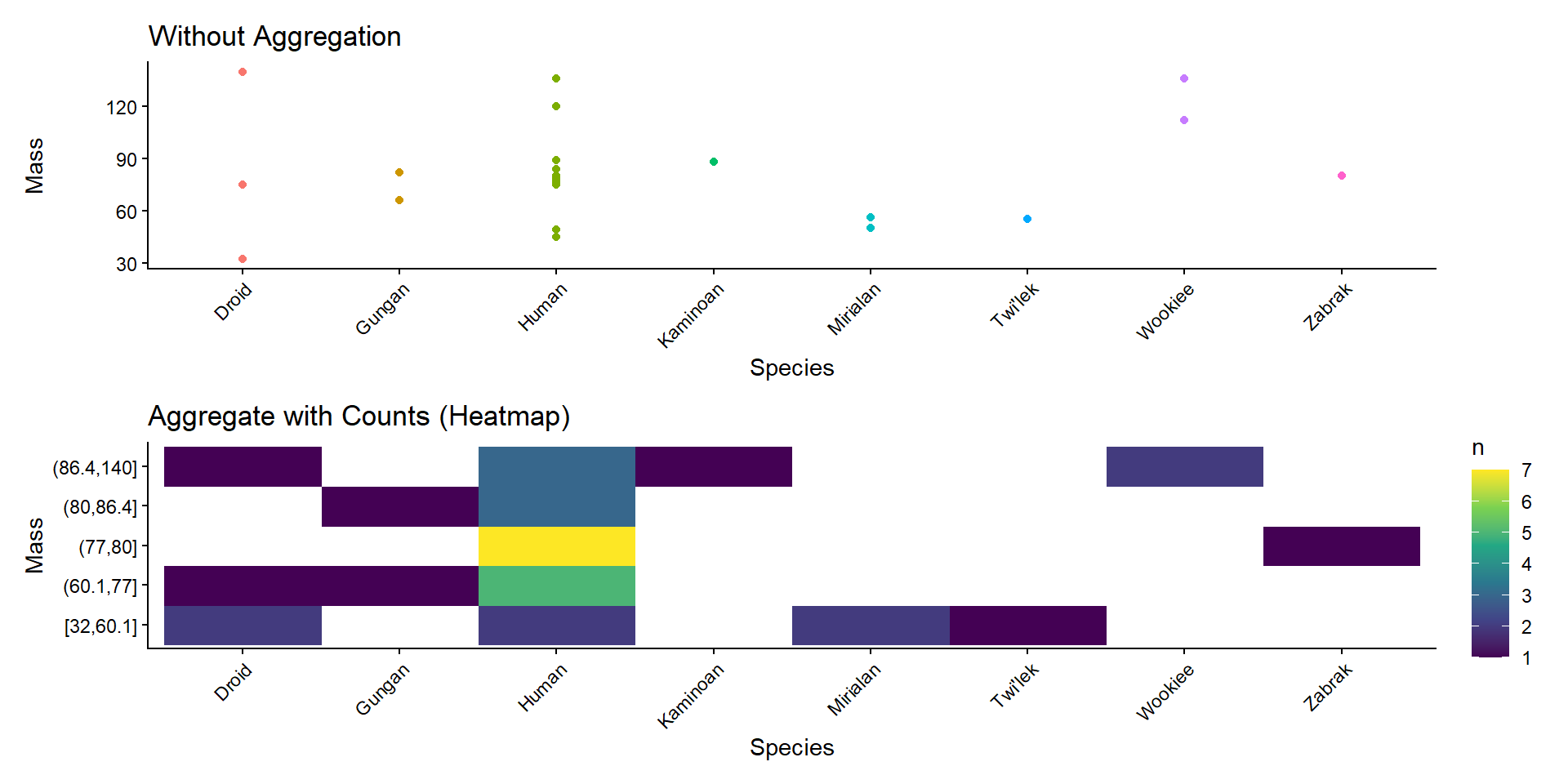
Item Aggregation
2D Summary Stats
Code
# 'sw' created on an earlier slide
a <- sw %>%
ggplot(aes(x = species, y = mass, color = species)) +
geom_point(show.legend = FALSE) +
ggtitle("Without Aggregation") +
labs(x = "Species", y = "Mass") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
b <- sw %>%
filter(!is.na(mass)) %>%
mutate(mass_bin = cut_number(mass, n = 5)) %>%
group_by(species, mass_bin) %>%
tally() %>%
ggplot(aes(x = species, y = mass_bin,
fill = n)) +
geom_raster() +
ggtitle("Aggregate with Counts (Heatmap)") +
labs(x = "Species", y = "Mass") +
scale_fill_viridis_c() +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
a/bAggregation
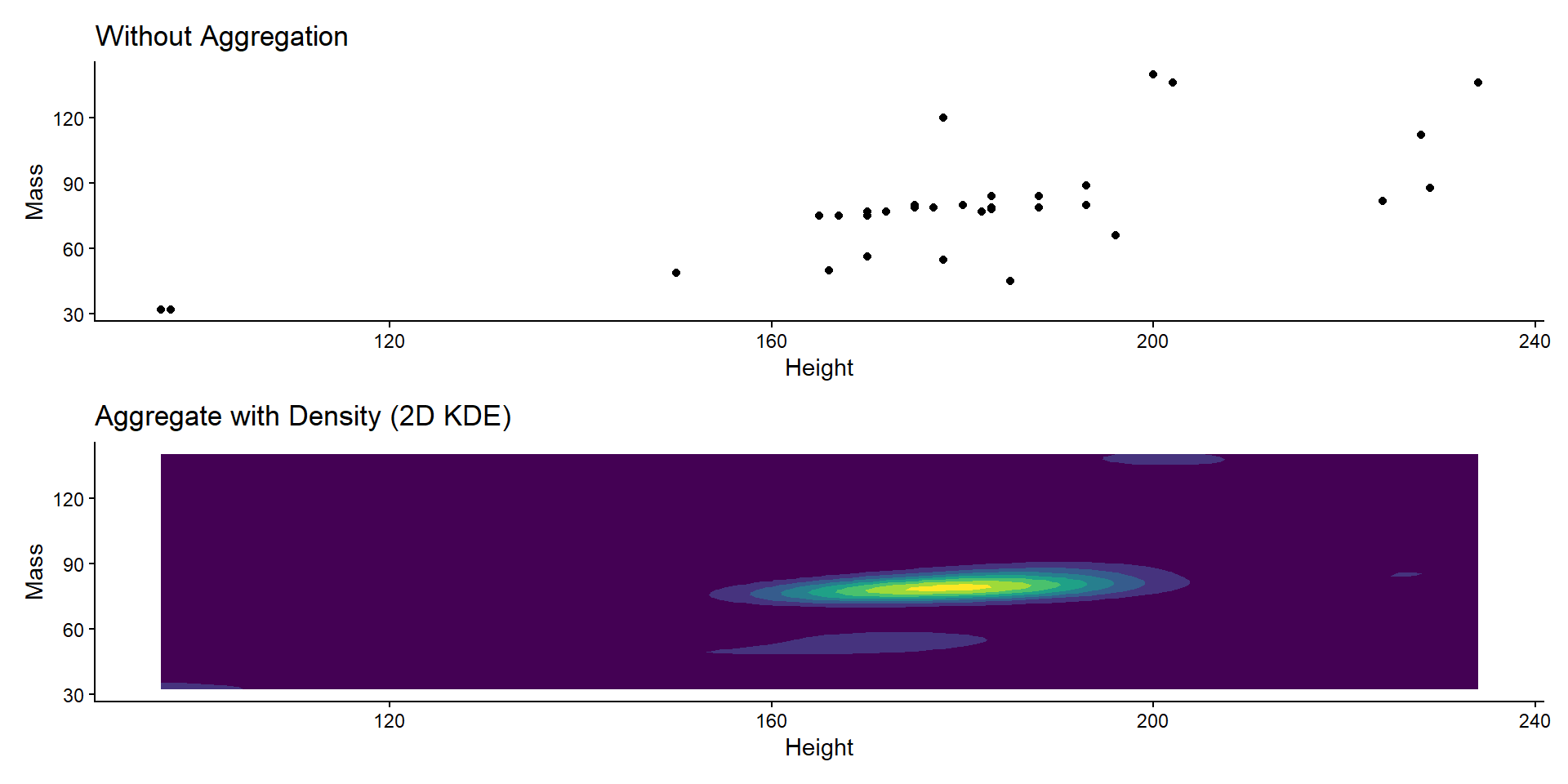
Item Aggregation
2D Summary Stats
Code
# 'sw' created on an earlier slide
a <- sw %>%
ggplot(aes(x = height, y = mass)) +
geom_point(show.legend = FALSE) +
ggtitle("Without Aggregation") +
labs(x = "Height", y = "Mass") +
theme_classic()
b <- sw %>%
filter(!is.na(mass)) %>%
ggplot(aes(x = height, y = mass)) +
geom_density_2d_filled(show.legend = FALSE) +
ggtitle("Aggregate with Density (2D KDE)") +
labs(x = "Height", y = "Mass") +
scale_fill_viridis_d() +
theme_classic()
a/bAggregation
Attribute Aggregation
- Generally refered to as dimensionality reduction (Wikipedia).
- Goal is preserve meaningful structure while using fewer attributes.
- Sometimes, simple solutions exist.
- Mostly involves complex mathematics.
Aggregation
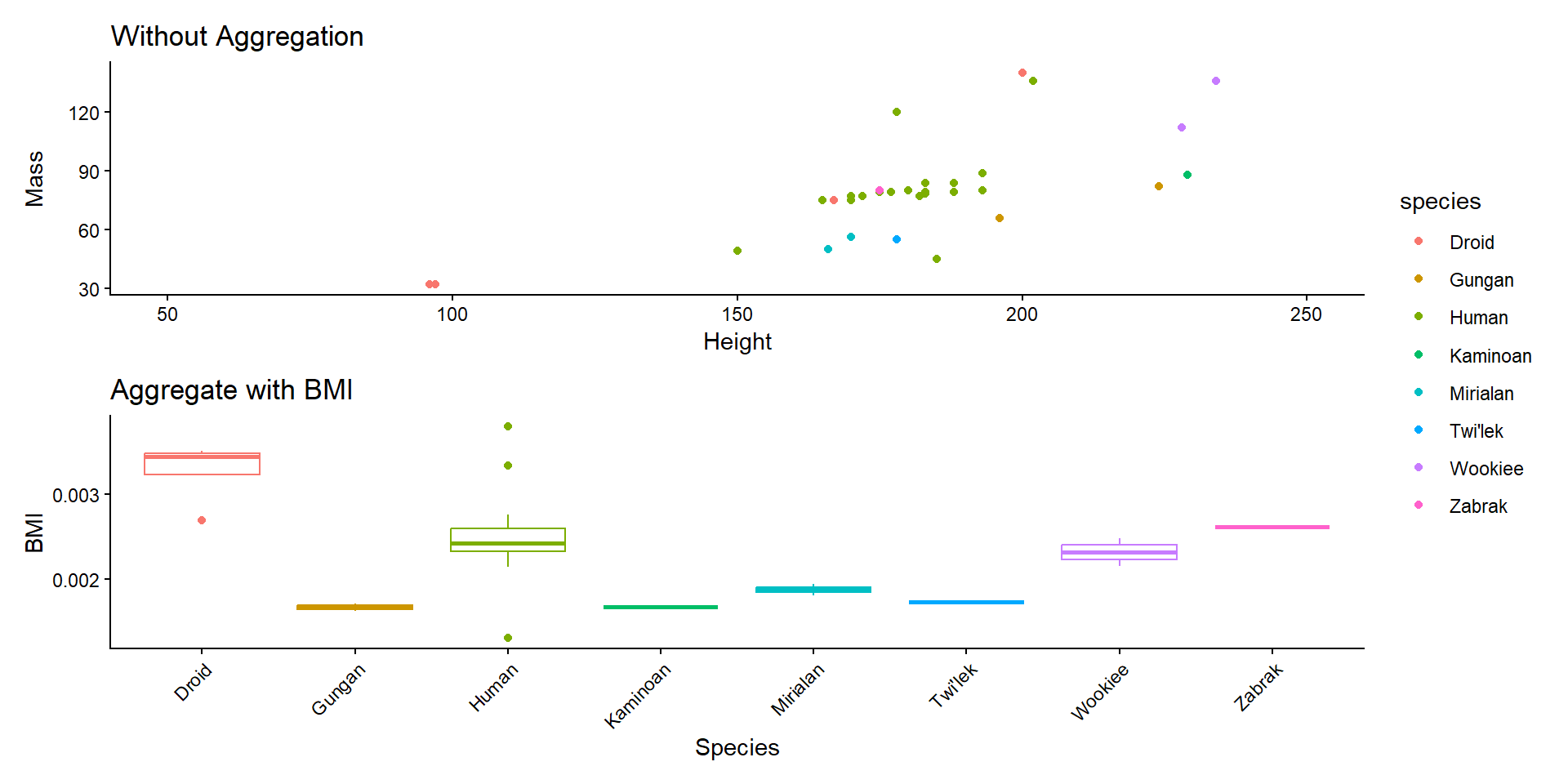
Attribute Aggregation
- Simple solution:
\[BMI = kg/m^2\]
Code
# 'sw' created on previous slide
a <- sw %>%
ggplot(aes(x = height, y = mass, color = species)) +
geom_point() +
coord_cartesian(xlim = c(50, 250)) +
ggtitle("Without Aggregation") +
labs(x = "Height", y = "Mass") +
theme_classic()
b <- sw %>%
mutate(bmi = mass/(height^2)) %>%
ggplot(aes(x = species, y = bmi,
color = species)) +
geom_boxplot(show.legend = FALSE) +
ggtitle("Aggregate with BMI") +
labs(x = "Species", y = "BMI") +
theme_classic() +
theme(axis.text.x = element_text(angle = 45, hjust = 1))
a / b + plot_layout(guides = "collect")Aggregation
Attribute Aggregation
Aggregation
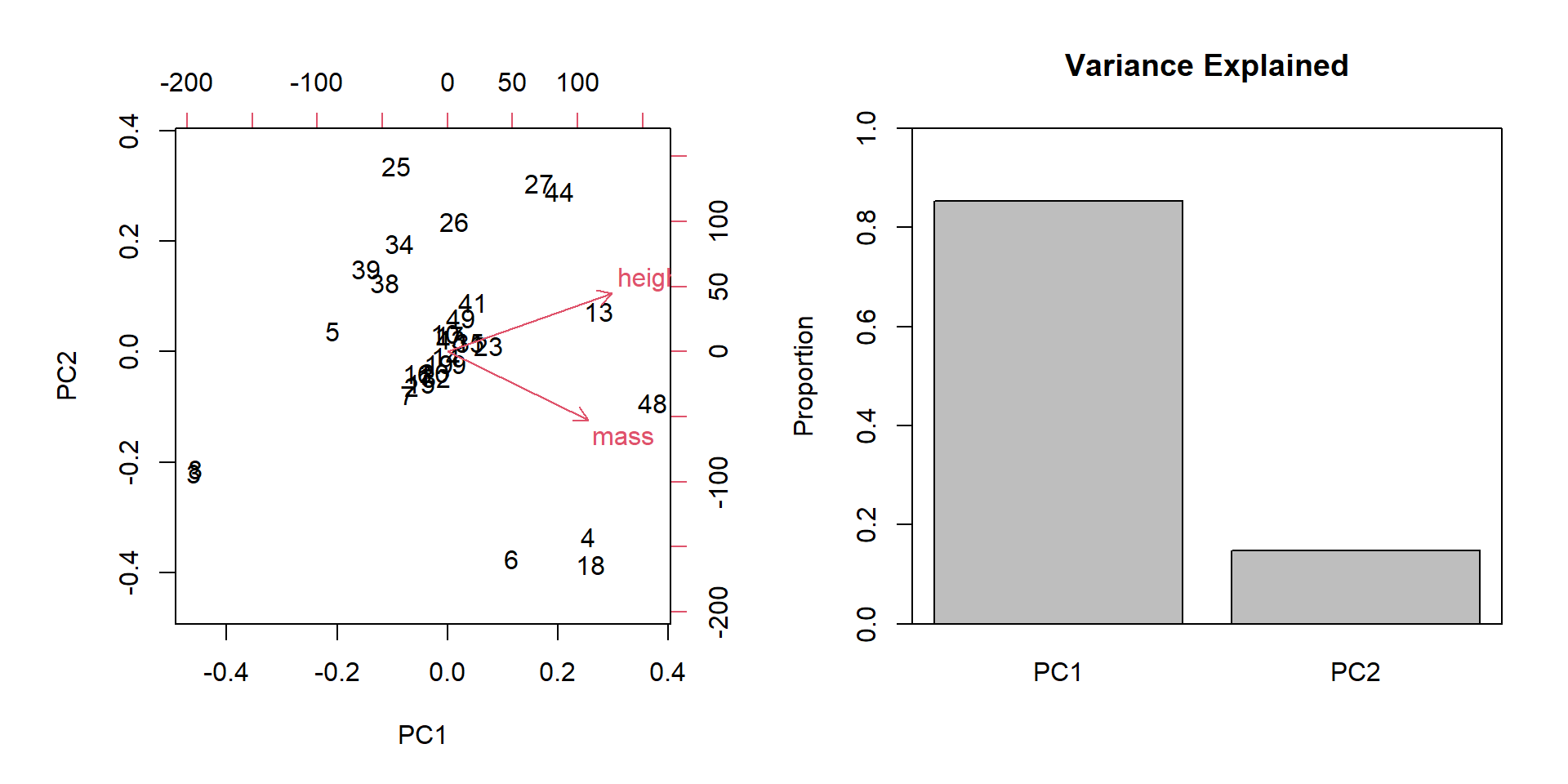
Attribute Aggregation
PCA
Code
# 'sw' created earlier; adding BMI here
sw <- sw %>%
mutate(bmi = mass/height^2)
pca <- prcomp(~ height + mass, data = sw)
pred <- predict(pca, sw)
sw$pc1 <- pred[, 1]
par(mfrow = c(1, 2))
biplot(pca)
summ <- summary(pca)
barplot(summ$importance[2, ],
ylim = c(0, 1), ylab = "Proportion",
main = "Variance Explained")
box()Aggregation
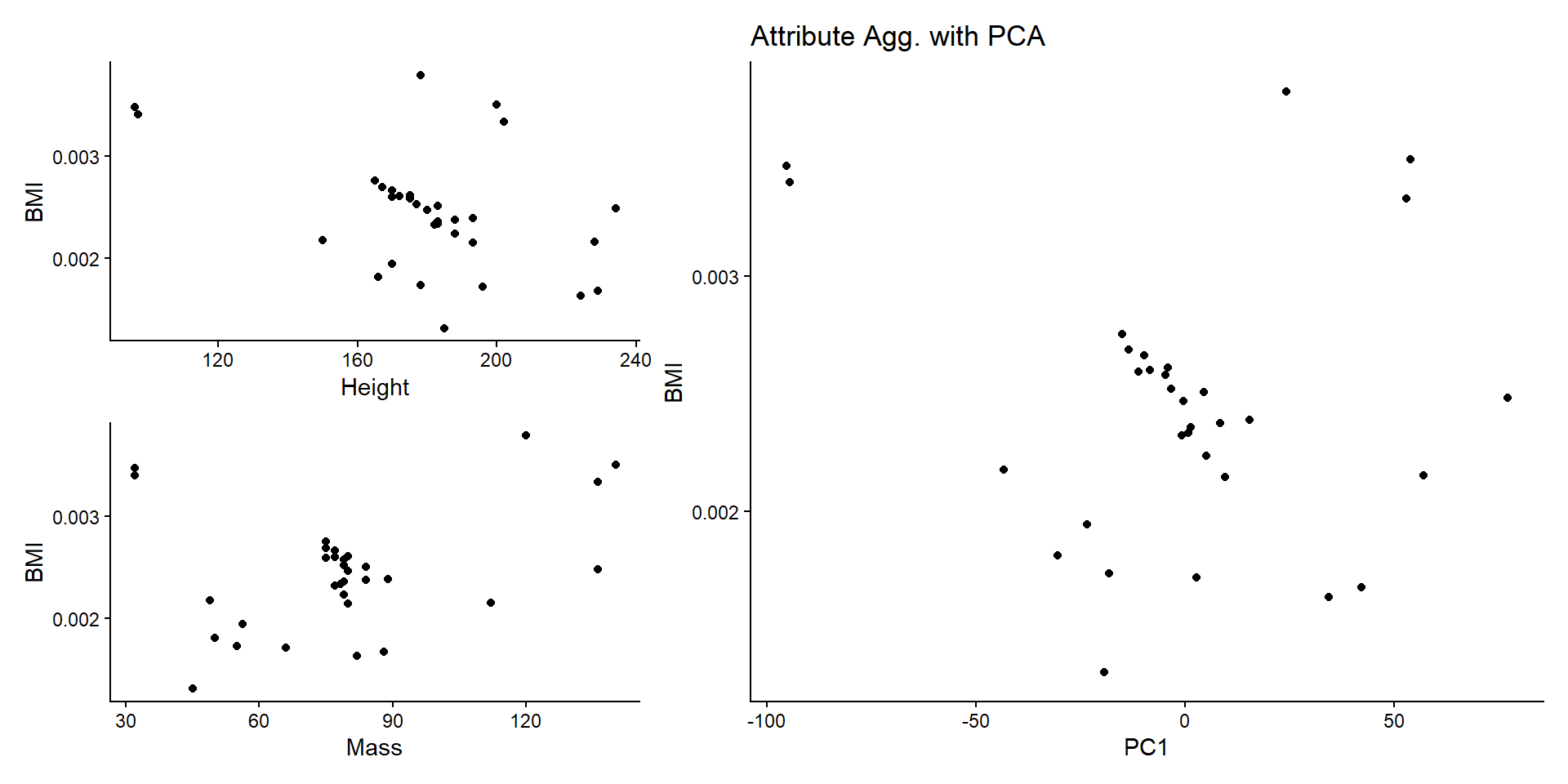
Attribute Aggregation
PCA
Code
# 'sw' created earlier
a <- sw %>%
ggplot(aes(x = height, y = bmi)) +
geom_point() +
labs(x = "Height", y = "BMI") +
theme_classic()
b <- sw %>%
ggplot(aes(x = mass, y = bmi)) +
geom_point() +
labs(x = "Mass", y = "BMI") +
theme_classic()
c <- sw %>%
ggplot(aes(x = pc1, y = bmi)) +
geom_point() +
labs(x = "PC1", y = "BMI") +
ggtitle("Attribute Agg. with PCA") +
theme_classic()
des <- "AC
BC"
a + b + c + plot_layout(widths = c(0.4, 0.6),
design = des)