# Create a graph
g1 <- make_graph(edges = c(1, 2, 2, 3, 3, 1,
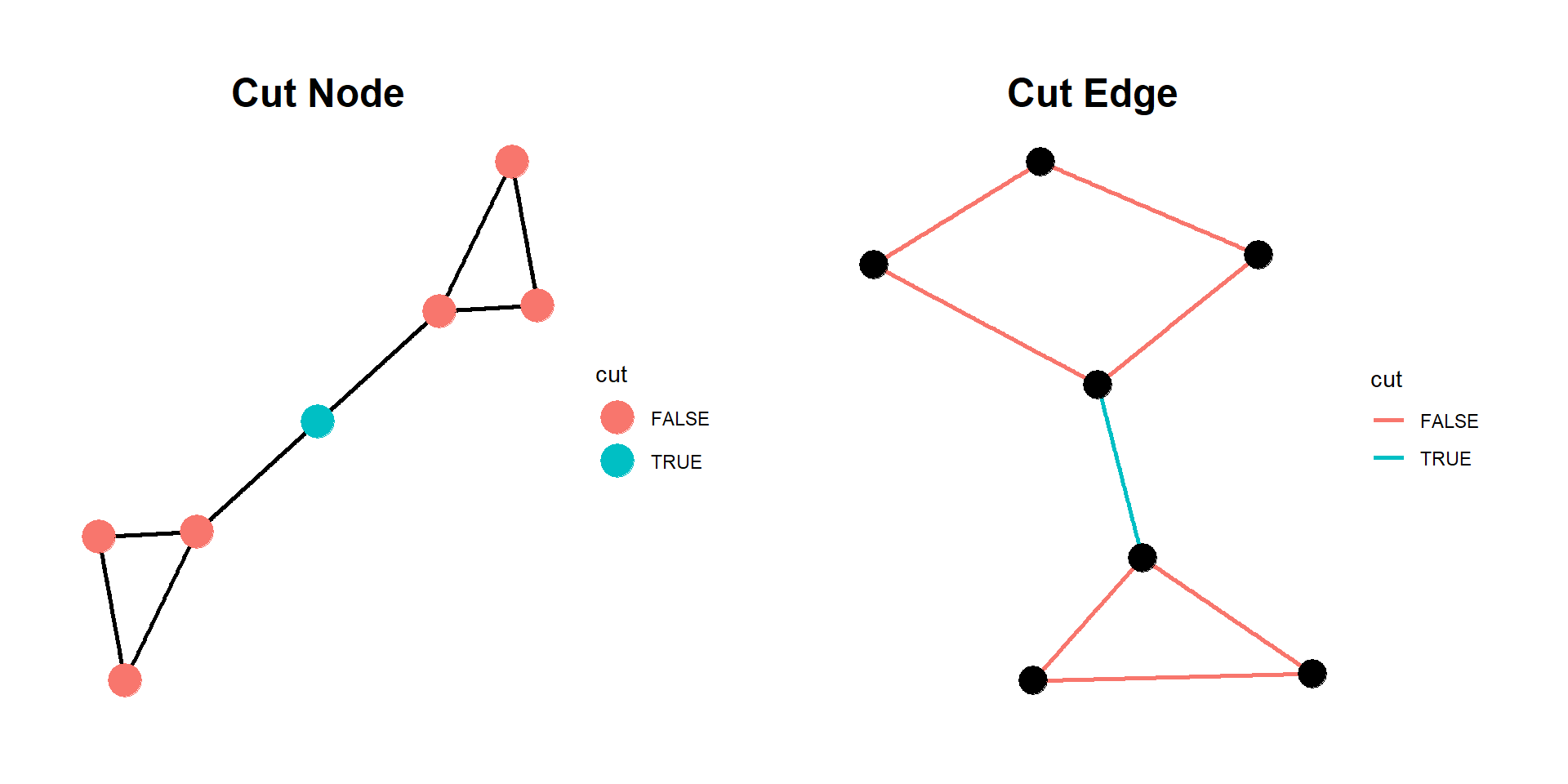
# Node 4 is cut node
3, 4, 4, 5,
5, 6, 6, 7, 7, 5),
directed = FALSE) %>%
set_vertex_attr(name = "cut", value = c((1:7) == 4))
g2 <- make_graph(edges = c(1, 2, 2, 3, 3, 1,
# Cut vertex between 3 and 4
3, 4,
4, 5, 5, 6, 6, 7, 7, 4),
directed = FALSE) %>%
set_edge_attr(name = "cut", value = c((1:8) == 4))
p1 <- ggraph(g1, layout = "igraph", algorith = "fr") +
geom_edge_link(linewidth = 1,
color = "black") +
geom_node_point(size = 7, aes(color = cut)) +
ggtitle("Cut Node") +
theme_graph() +
theme(plot.title = element_text(hjust = 0.5))
p2 <- ggraph(g2, layout = "igraph", algorith = "fr") +
geom_edge_link(linewidth = 1, aes(color = cut)) +
geom_node_point(size = 6) +
ggtitle("Cut Edge") +
theme_graph() +
theme(plot.title = element_text(hjust = 0.5))
p1 + p2